SleepFM: AI Predicts 130 Diseases From One Night of Sleep
Authors: Rahul Thapa, Magnus Ruud Kjaer, Emmanuel Mignot, James Zou
Every night you broadcast. Your brain fires electrical patterns across a dozen electrodes. Your heart drums a rhythm that subtly shifts between sleep stages. Your diaphragm contracts 6,000 times. Your legs twitch. Your eyes dart. Seven streams of physiological data, eight hours straight — and after a sleep technician scores the stages, nearly all of it goes to waste.
A team at Stanford decided to stop wasting it. They fed 585,000 hours of overnight sleep recordings from 65,000 patients into a neural network and published the results in Nature Medicine in January 2026. The model, called SleepFM, learned to predict 130 medical conditions from a single night’s recording. Parkinson’s disease, with a concordance index of 0.89. Breast cancer, 0.87. Dementia, 0.85. Years before the first clinical symptom.
The Largest Sleep Dataset Ever Fed to a Machine
Polysomnography (PSG) is the gold standard of sleep studies. A patient spends one night in a lab wired with sensors on the scalp, chest, legs, and fingertip. The system records: electroencephalography (EEG), electrocardiography (ECG), electromyography (EMG), eye movements, airflow, blood oxygen saturation, and leg movements.
The core of the dataset came from 35,000 patients at the Stanford Sleep Medicine Center, followed for up to 25 years after their overnight lab visit — long enough to know who developed disease and who did not. On top of that: clinical recordings from Denmark and France, plus three major epidemiological cohorts (MESA, MrOS, and SHHS). In total, 65,649 people aged 2 to 96, generating 585,000 cumulative hours of continuous physiological data.

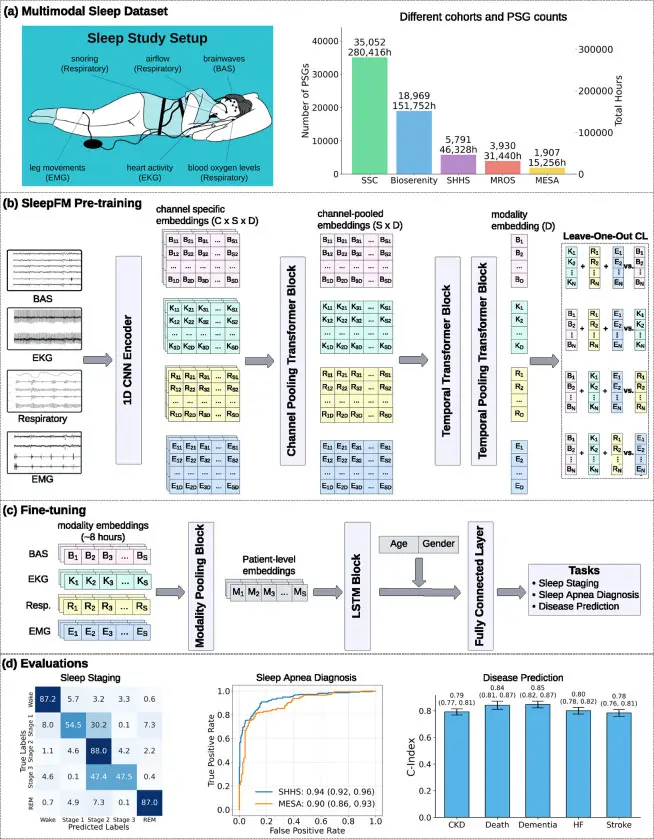
Fig. 1: SleepFM architecture overview. Left: data sources and signal types. Center: multimodal contrastive learning. Right: fine-tuning for classification and disease prediction. Source: Thapa et al., Nature Medicine, 2026
How a Neural Network Learns to Read Sleep
«SleepFM is essentially learning the language of sleep, ” says James Zou, associate professor of biomedical data science at Stanford and one of the study’s senior authors. The metaphor is surprisingly literal.
Foundation model — a neural network trained on massive data without a specific task in mind. Just as GPT learns language by reading billions of words, SleepFM learns the structure of sleep by processing hundreds of thousands of nights.
The model chops each recording into five-second tokens — sleep’s equivalent of words. Six convolutional layers encode the raw signal at 128 Hz, then a transformer with eight attention heads merges information across channels. The whole thing weighs 4.44 million parameters — tiny by language model standards, but enough to capture the body’s nocturnal vocabulary.
The training trick is called leave-one-out contrastive learning. Hide the ECG channel and force the network to reconstruct cardiac rhythm from brainwaves and breathing alone. Hide the EEG and make it infer brain activity from everything else. This teaches SleepFM the deep, cross-system correlations between organs — precisely the correlations that carry disease signals.
Pre-training took 15 hours on a single NVIDIA A100 GPU. Fine-tuning for a specific condition: two to five minutes.
Parkinson’s, Seven Years Before the Tremor Starts
The researchers tested SleepFM against 1,041 medical diagnoses coded in patient records. Of those, 130 showed statistically significant predictive performance with a concordance index at or above 0.75 — the threshold where a prognostic model becomes clinically informative.
C-index (concordance index) measures how well a model ranks patients by risk over time. A score of 0.5 is random guessing. A score of 1.0 is perfect. A C-index of 0.89 for Parkinson’s means: given any two patients, the model correctly identified which one would develop the disease first in 89% of pairs.
The numbers hold up across disease categories. Parkinson’s disease: 0.89. Prostate cancer: 0.89. Breast cancer: 0.87. Dementia: 0.85. Hypertensive heart disease: 0.84. All-cause mortality: 0.84. Myocardial infarction: 0.81. Mild cognitive impairment: 0.81.
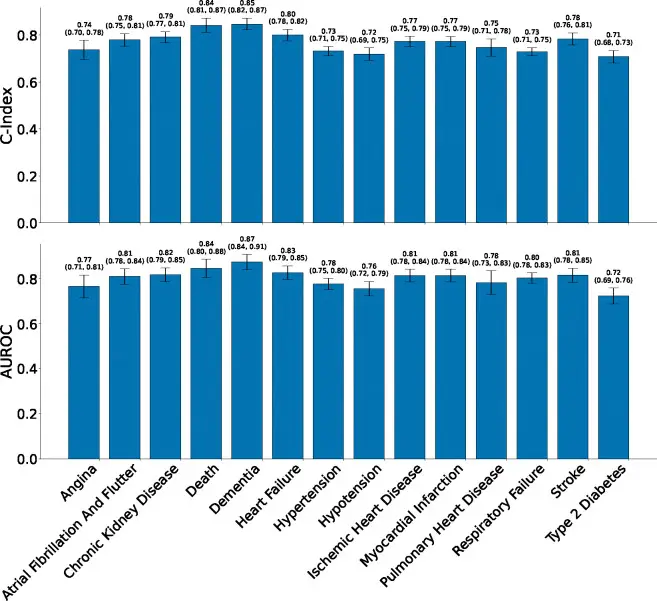
Fig. 2: Predictive performance for 14 key conditions. SleepFM (red) consistently outperforms both the demographics-only model (gray) and the non-pretrained baseline (blue). Source: Thapa et al., Nature Medicine, 2026
A demographics-only model using age, sex, BMI, and ethnicity performed substantially worse — confirming that SleepFM extracts genuine sleep-encoded disease information beyond what a basic risk calculator provides.
What Each Sleep Stage Whispers About Your Health
The most unexpected finding was not what SleepFM predicts, but where the signal lives. The team decomposed predictions by sleep stage and by recording channel.
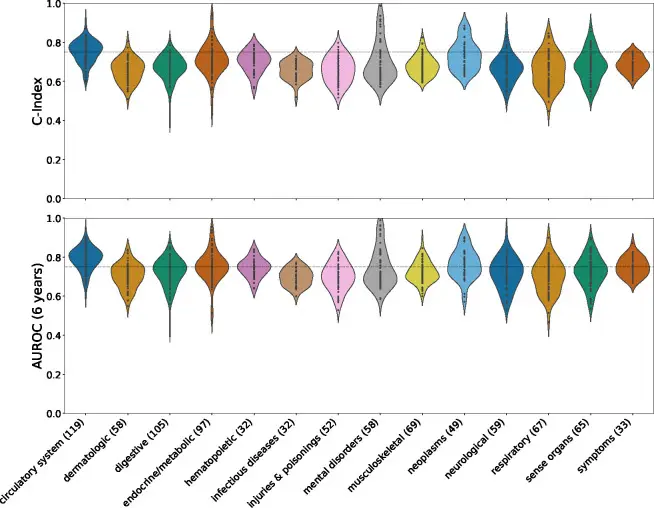
Fig. 3: Heatmap showing C-index and 6-year AUROC across major disease categories. Source: Thapa et al., Nature Medicine, 2026
REM sleep emerged as the strongest predictor for dementia, chronic kidney disease, and respiratory failure. Light sleep (stages N1 and N2) dominated predictions for heart failure, stroke, angina, and type 2 diabetes with renal complications. Deep sleep (N3), the stage most people associate with restorative rest, turned out to be the weakest predictor — only 2 of the 62 best-performing conditions relied primarily on N3.
The channel breakdown was equally telling. ECG led for 47 conditions — mostly cardiovascular. Brain signals (EEG) dominated 32 neurological diagnoses: autism, dementia, Parkinson’s, developmental delays. Respiratory channels contributed to another 32 conditions, with an unexpected guest among them: melanoma. The link between breathing patterns during sleep and a systemic cancer remains unexplained and will require dedicated follow-up studies.
«The most information we got for predicting disease was by contrasting the different channels, ” said Emmanuel Mignot, the world’s leading narcolepsy expert and the study’s co-senior author.
The Gap Between a Paper and a Patient’s Bedside
SleepFM is a rigorous piece of science, but a foundation model in Nature Medicine is not a clinical tool. The study was published in Nature Medicine on January 6, 2026, and has undergone peer review, though independent replication of disease-prediction results on fully external cohorts remains limited.
Generalizability is the first concern. A third of the training data came from Stanford’s own sleep clinic. Patients referred to sleep labs are not a random sample of the population — they already have symptoms worth investigating. The authors validated SleepFM on the SHHS cohort (completely excluded from training), and the model maintained predictive ability for stroke and cardiovascular mortality. But under a temporal shift — training on pre-2020 data, testing on post-2020 patients — accuracy dropped noticeably. Models age alongside the equipment that recorded their inputs.
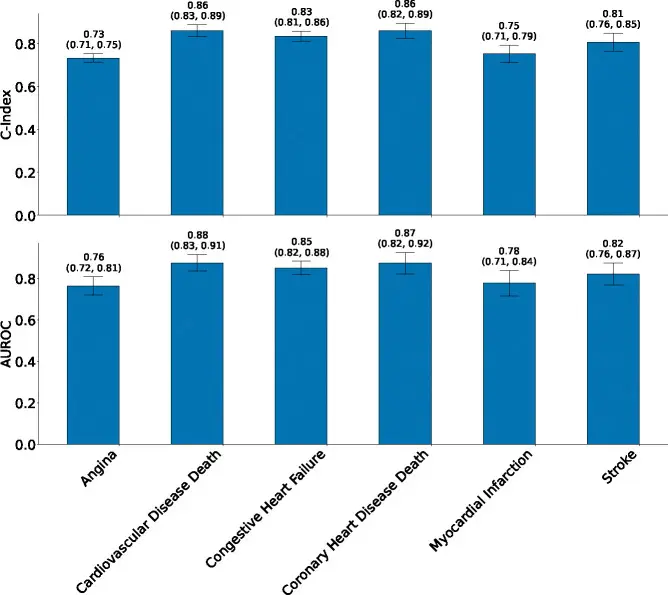
Fig. 4: Transfer validation on the SHHS cohort, held out entirely from pre-training. SleepFM maintains predictive performance for stroke and cardiovascular mortality. Source: Thapa et al., Nature Medicine, 2026
Interpretability is the second. The model produces numbers, not explanations. «It doesn’t explain that to us in English, ” Zou admits. For a physician who must justify a risk score to a patient, that opacity matters. The team is developing separate interpretation tools, but they remain partial.
The third gap is outcomes. Predicting risk is not the same as reducing it. No prospective trial has tested whether alerting patients based on SleepFM outputs improves health outcomes — and without one, clinical utility remains theoretical. Regulatory approval from the FDA or EMA is not on the immediate horizon.
From Sleep Labs to Wrist Sensors
Despite its limitations, SleepFM proved something fundamental: a single night of sleep encodes clinically meaningful information about dozens of conditions unrelated to sleep disorders. Not a screening for apnea — a full-body diagnostic map, encrypted in the rhythms of the night.
The model’s code is open-source on GitHub. The team plans two next steps: integrating wearable sensor data (smartwatches, sleep rings) and building interpretation tools for clinicians. If SleepFM or its successors learn to work with simplified signals — one or two channels from a wrist device instead of seven from a lab — the clinical implications would be enormous.
Every night, your body transmits hundreds of thousands of signals. For the first time, there is a system designed to listen.
Frequently Asked Questions
Can I take the SleepFM test at home with a smartwatch?
Not yet. SleepFM was trained on clinical polysomnography data — seven sensor types at 128 Hz sampling rate. Consumer devices like Apple Watch or Oura Ring capture far fewer channels with lower resolution. The authors plan to adapt the model for simplified signals, but that adaptation is a separate research effort.
How accurate are SleepFM’s predictions for an individual person?
A C-index of 0.89 for Parkinson’s means excellent group-level risk ranking, not individual diagnosis. The model identifies who in a population faces higher risk — it does not diagnose a specific patient. Think of it as a screening tool, not a replacement for an MRI or biopsy.
Which diseases does SleepFM predict best?
The highest concordance indices were achieved for Parkinson’s disease (0.89), prostate cancer (0.89), breast cancer (0.87), dementia (0.85), hypertensive heart disease (0.84), and all-cause mortality (0.84). In total, 130 diagnoses showed statistically significant predictive ability above the 0.75 threshold.
Why was deep sleep the weakest predictor?
The authors suggest that N3 (deep sleep) contains less variable patterns across patients. REM and light sleep stages show greater inter-individual differences and therefore carry more diagnostic information. Deep sleep is dominated by slow waves, which appear less sensitive to systemic disease signatures.
When might SleepFM reach a regular clinic?
Not soon. The model needs independent external validation across diverse countries and ethnic groups, prospective clinical trials, and regulatory approval (FDA/EMA). The open-source code on GitHub enables other teams to begin validation now — and that is likely the fastest route to the clinic.
References
Related
Context
Related Articles
How Sleep Loss Turns Your Gut Against Your Brain
Gut bacteria from sleep-deprived mice triggered Alzheimer's-like tau damage in healthy brains. Scientists traced the full molecular chain.
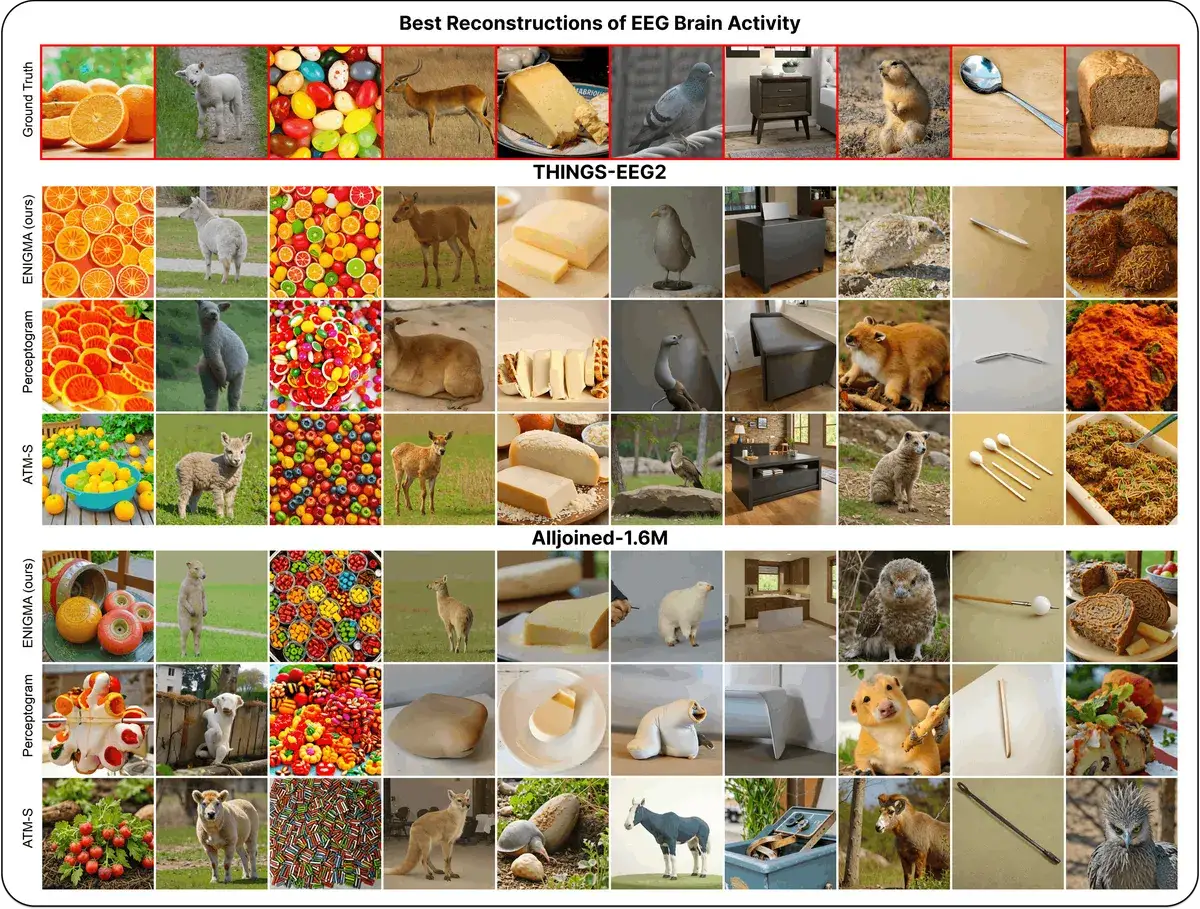
ENIGMA: How to Read Minds in 15 Minutes with a $2,200 Headset
ENIGMA reconstructs images from brain signals (EEG) after just 15 minutes of calibration, using less than 1% of previous methods' parameters. Works even with consumer-grade headsets.
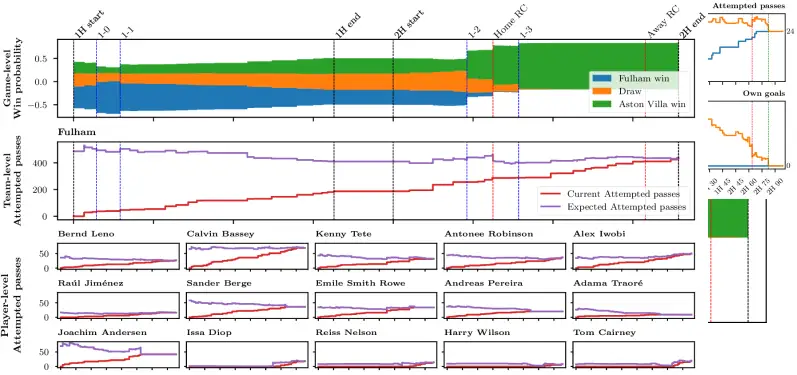
75,000 Predictions Per Match: Axial Transformer Forecasts Live Football
Stats Perform researchers built an Axial Transformer neural network that generates 75,000 live predictions per football match with sub-second latency